 If you have never installed it before you can also use the install.packages("tidyverse") call to install it for the first time. Using the library function we will load the tidyverse. You could think of it like being able to write R code inside a basic Word document (but it can do a lot more than that!).Īlthough you may not be interested in the dataset I have provided, this hopefully provides a clear workflow for you to swap in your data of interest and accomplish a basic analysis! Load the tidyverse, broom, and knitr R Markdown is a document created inside R that allows you to write code, execute it inline, and write comments/notes as you go. If you would rather see the entire workflow in an R-Markdown document, please see here. You can simply copy-paste the code seen here and it will run in R. I will use knitr::kable to generate some html tables for a markdown document, but it is not necessary for the workflow.Īdditionally, I will be uploading the Excel Sheet used in this example, so that you can re-create the workflow on your own. While other stats-heavy packages provide additional statistical testing, base R has a decent ability to perform statistical analyses out of the box. 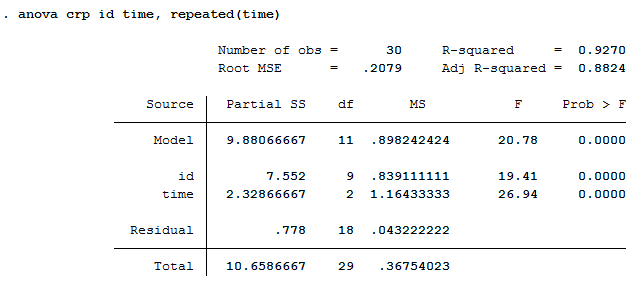
These two packages dramatically improve the data analysis workflow in my opinion. We will limit dependence to two packages: tidyverse and broomwhile using base R for the rest. We will be working through loading, plotting, analyzing, and saving the outputs of our analysis through the tidyverse, an “opinionated collection of R packages” designed for data analysis. That’s all! Please comment below if you have any questions.Source: Folstein et al, 1975, J Psychiatr Res 12:189–198 Another thing that you have to alter is selecting the appropriate range of column names that may differ in your data set. You can choose the one that best describe your experimental design. Here are some of the model terms for different experimental designs. Then you need to select a model term that best describes the experimental design. You must provide the names of factor variables from your data set at the beginning part when we transformed variables from character to factor. Do you believe it’s a matter of a few clicks to produce a publication-ready ANOVA table in R after such tedious work? The answer is yes you can get the same output by just altering few parameters for your own data. This is how the ANOVA table will appear in the end. You may choose from a variety of document types. By using the part parameter as header in the bold() function, you may bold the column names.Ĭlick the compile report icon to produce this table in several document formats. We will supply the aov.table object in the data parameter, and we may also give the column name for the table’s rownames following the pipe operator. To do so, we’ll utilise the flextable function from the flextable package. Load these packages by using library() function.įinally, in an MS Word document, we must print this publication-ready ANOVA table. The packages that must be loaded before utilizing certain functions are listed here. This has a significant benefit over typical manual procedures for computation and table display, which are not only time-consuming but also prone to errors. This will help researchers to quickly create ANOVA tables while saving time by eliminating the need to manually input each command. None of the statistical software can generate such tables without advanced knowledge of the corresponding programming language. You will be able to export this table in MS word, PDF and HTML document. 
Here, we shall focus on introducing a new method capable of generating publication ready ANOVA tables within few simple steps. One of the most important statistical tables is Analysis of variance (ANOVA) and may be found in a wide range of agricultural, biological and medicinal research. Traditional manual procedures for computing and presenting statistically significant findings are time-consuming and error-prone. In the biological sciences, statistical tables are an important part of data analysis and reporting. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
